Developing Methodology for Automatic Generation of Evolvable Genotype-Phenotype Maps
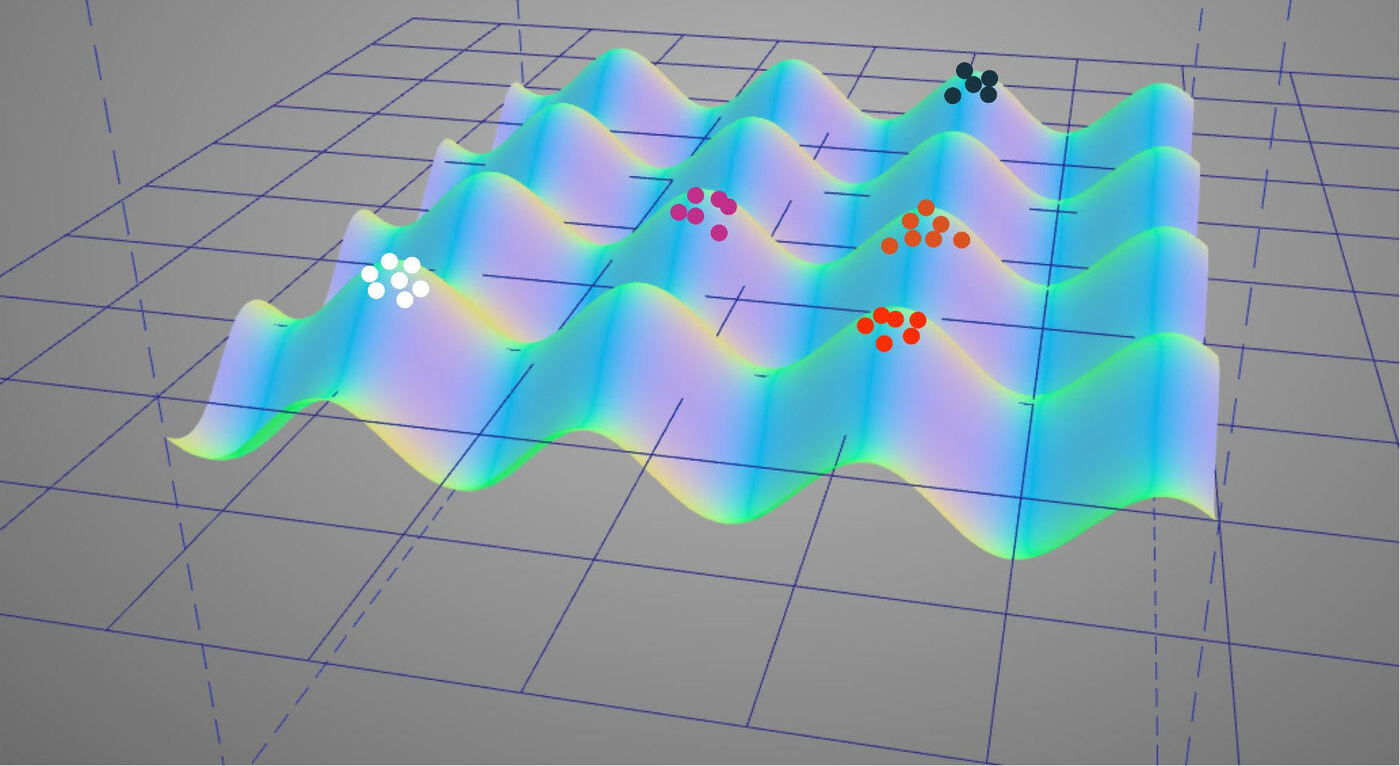 Visualization of rugged fitness landscape suited to denoising autoencoder genotype-phenotype map.
Visualization of rugged fitness landscape suited to denoising autoencoder genotype-phenotype map.
In biology, phenotype refers to an organism’s observable characteristics (morphological, behavioral, physiological, chemical, etc.). Importantly, the phenotype governs an organism’s ability to survive and reproduce — its fitness. Likewise, genotype refers to the heritable information that shapes an organism’s phenotype. This is typically equated with an organism’s DNA content.
The genotype-phenotype map describes the relationship between an organism’s genotype and its phenotype. In biology, this concept is tightly entwined with the process of development, the dynamics through which an organism’s genotype and environment interact to determine its phenotype. As might be expected, the genotype-phenotype map profoundly influences the character of phenotypic variation that is produced under genetic mutation.
Evolutionary search cannot succeed without the production of heritable, viable phenotypic variation, a trait referred to as evolvability. How to engineer genotype-phenotype maps to promote evolvability in digital evolution systems remains a crucial open question.
This work introduces the use of bottlenecked and denoising autoencoders to automatically generate evolvable genotype-phenotype mappings. Autoencoders are algorithms that learn to regurgitate a particular type of input, usually implemented using deep learning. They allow for enhancement of evolvability due to their ability to repair poor-quality solutions and compactly represent the space of high-quality solutions.
Publications & Software
View at Publisher
| Authors | Matthew Andres Moreno, Wolfgang Banzhaf, Charles Ofria |
| Date | July 15th, 2018 |
| DOI | 10.1145/3205455.3205597 |
| Venue | The Genetic and Evolutionary Computation Conference |
Abstract
We present AutoMap, a pair of methods for automatic generation of evolvable genotype-phenotype mappings. Both use an artificial neural network autoencoder trained on phenotypes harvested from fitness peaks as the basis for a genotype-phenotype mapping. In the first, the decoder segment of a bottlenecked autoencoder serves as the genotype-phenotype mapping. In the second, a denoising autoencoder serves as the genotype-phenotype mapping. Automatic generation of evolvable genotype-phenotype mappings are demonstrated on the n-legged table problem, a toy problem that defines a simple rugged fitness landscape, and the Scrabble string problem, a more complicated problem that serves as a rough model for linear genetic programming. For both problems, the automatically generated genotype-phenotype mappings are found to enhance evolvability.
BibTeX
@inproceedings{moreno2018learning,
author = {Moreno, Matthew Andres and Banzhaf, Wolfgang and Ofria, Charles},
title = {Learning an Evolvable Genotype-Phenotype Mapping},
year = {2018},
isbn = {9781450356183},
publisher = {Association for Computing Machinery},
address = {New York, NY, USA},
url = {https://doi.org/10.1145/3205455.3205597},
doi = {10.1145/3205455.3205597},
booktitle = {Proceedings of the Genetic and Evolutionary Computation Conference},
pages = {983–990},
numpages = {8},
keywords = {deep learning, indirect encodings, evolvability, genetic algorithms, adaptive representations, genotype-phenotype map},
location = {Kyoto, Japan},
series = {GECCO '18}
}
Citation
Matthew Andres Moreno, Wolfgang Banzhaf, and Charles Ofria. 2018. Learning an evolvable genotype-phenotype mapping. In Proceedings of the Genetic and Evolutionary Computation Conference (GECCO ‘18). Association for Computing Machinery, New York, NY, USA, 983–990. https://doi.org/10.1145/3205455.3205597